Making Life Better
through
Education, Research & Patient Care.
Pause Video
USF HEALTH
USF HEALTH
Education
Training the next generation of health care professionals to improve patient safety and health outcomes.
Research
Accelerating discoveries into innovative treatments, diagnostics and devices for complex medical conditions.
Patient Care
Providing high-quality care as the region's only academic medical center with more than 1,000 providers and 30 specialties.
Student Success
As part of an academic health center within a Preeminent Research University, USF Health is dedicated to the academic success of more than 6,000 students annually who will be workforce ready in medicine, nursing, public health, pharmacy, and physical therapy. Students work together as teams with access to the latest knowledge, advanced research, and real-world simulation-based training to transform health care in our region, state, and nation.
Learn More
USF Health in
Downtown Tampa
Our USF Health building in downtown Tampa is a 395,000-square-foot, state-of-the-art facility featuring world-class labs, high-tech learning areas, and top-notch research spaces. It houses the USF Health Morsani College of Medicine, Taneja College of Pharmacy, and the Heart Institute. Situated near Tampa General Hospital, the USF Health South Tampa Center for Advanced Healthcare, and the USF Health Center for Advanced Medical Learning and Simulation (CAMLS), our iconic building is transforming education, research, and patient care in our region and beyond.
Together with Tampa General Hospital, we have created the Tampa Medical and Research District, a life sciences epicenter in downtown Tampa. This hub attracts top talent and innovation, driving job creation, regional growth, and enhancing access to cutting-edge research and education for patients and students.
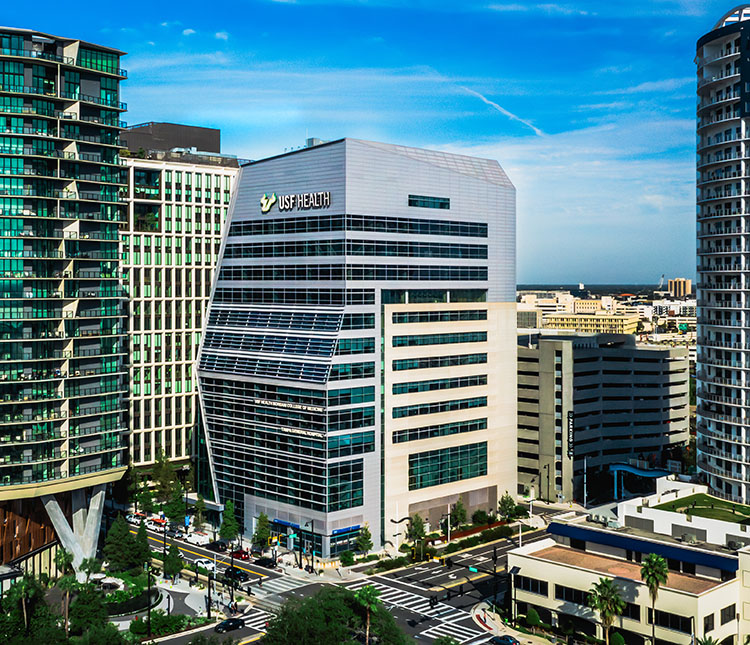

Downtown Tampa
Morsani College of Medicine, Taneja College of Pharmacy and Heart Institute.



Delivering Health Excellence
Charles J. Lockwood, MD, MHCM, executive vice president of USF Health and dean of the USF Health Morsani College of Medicine, shares his thoughts on academic medicine and health news.
Learn MoreUSF Health News

USF Health & St. Pete College PT/PTA Students Team Up for IPE Day to Enhance Patient Care & Teamwork
USF Health and St. Petersburg College students from Physical Therapy (PT) and Physical Therapist Ass...

Three USF Health faculty among 2024 class of Fellows of American Association for the Advancement of Science
11 USF faculty were inducted, marking the third largest cohort from any university in the nation. Re...

Class of 2025 medical students celebrate ‘magical’ Match Day
Class of 2025 medical students celebrate "magical" Match Day and learn where they will spend their r...